PRISM includes a (prototype) tool to translate specifications in SBML (Systems Biology Markup Language) to model descriptions in the PRISM language. SBML is an XML-based format for representing models of biochemical reaction networks. The translator currently works with Level 2 Version 1 of the SBML specification, details of which can be found here. Level 1 SBML files will first need to be translated into equivalent Level 2 files, for example using this on-line converter.
Since PRISM is a tool for analysing discrete-state systems, the translator is designed for SBML files intended for discrete stochastic simulation. A useful set of such files can be found in the CaliBayes Discrete Stochastic Model Test Suite. There are also many more SBML files available in the BioModels Database.
We first give a simple example of an SBML file and its PRISM translation. We then give some more precise details of the translation process.
Example
An SBML file comprises a set of species and a set of reactions which they undergo. Below is the SBML file for the simple reversible reaction: Na + Cl ↔ Na+ + Cl-, where there are initially 100 Na and Cl atoms and no ions, and the base rates for the forwards and backwards reactions are 100 and 10, respectively.
<?xml version="1.0" encoding="UTF-8"?>
<sbml xmlns="http://www.sbml.org/sbml/level2" metaid="_000000" level="2" version="1">
<model id="nacl" name="Na+Cl">
<listOfCompartments>
<compartment id="compartment"/>
</listOfCompartments>
<listOfSpecies>
<species id="na" initialAmount="100" hasOnlySubstanceUnits="true"/>
<species id="cl" initialAmount="100" hasOnlySubstanceUnits="true"/>
<species id="na_plus" initialAmount="0" hasOnlySubstanceUnits="true"/>
<species id="cl_minus" initialAmount="0" hasOnlySubstanceUnits="true"/>
</listOfSpecies>
<listOfReactions>
<reaction id="forwards" reversible="false">
<listOfReactants>
<speciesReference species="na"/>
<speciesReference species="cl"/>
</listOfReactants>
<listOfProducts>
<speciesReference species="na_plus"/>
<speciesReference species="cl_minus"/>
</listOfProducts>
<kineticLaw>
<math xmlns="http://www.w3.org/1998/Math/MathML">
<apply><times/><ci>forwards_rate</ci>
<apply><times/><ci>na</ci><ci>cl</ci></apply></apply>
</math>
<listOfParameters>
<parameter id="forwards_rate" value="100"/>
</listOfParameters>
</kineticLaw>
</reaction>
<reaction id="backwards" reversible="false">
<listOfReactants>
<speciesReference species="na_plus"/>
<speciesReference species="cl_minus"/>
</listOfReactants>
<listOfProducts>
<speciesReference species="na"/>
<speciesReference species="cl"/>
</listOfProducts>
<kineticLaw>
<math xmlns="http://www.w3.org/1998/Math/MathML">
<apply><times/><ci>backwards_rate</ci>
<apply><times/><ci>na_plus</ci><ci>cl_minus</ci></apply></apply>
</math>
<listOfParameters>
<parameter id="backwards_rate" value="10"/>
</listOfParameters>
</kineticLaw>
</reaction>
</listOfReactions>
</model>
</sbml>
And here is the resulting PRISM code:
ctmc
const int MAX_AMOUNT = 100;
const double forwards_rate = 100;
const double backwards_rate = 10;
const int na_MAX = MAX_AMOUNT;
module na
na : [0..na_MAX] init 100;
[forwards] na > 0 -> (na'=na-1);
[backwards] na <= na_MAX-1 -> (na'=na+1);
endmodule
const int cl_MAX = MAX_AMOUNT;
module cl
cl : [0..cl_MAX] init 100;
[forwards] cl > 0 -> (cl'=cl-1);
[backwards] cl <= cl_MAX-1 -> (cl'=cl+1);
endmodule
const int na_plus_MAX = MAX_AMOUNT;
module na_plus
na_plus : [0..na_plus_MAX] init 0;
[forwards] na_plus <= na_plus_MAX-1 -> (na_plus'=na_plus+1);
[backwards] na_plus > 0 -> (na_plus'=na_plus-1);
endmodule
const int cl_minus_MAX = MAX_AMOUNT;
module cl_minus
cl_minus : [0..cl_minus_MAX] init 0;
[forwards] cl_minus <= cl_minus_MAX-1 -> (cl_minus'=cl_minus+1);
[backwards] cl_minus > 0 -> (cl_minus'=cl_minus-1);
endmodule
module reaction_rates
[forwards] (forwards_rate*(na*cl)) > 0 -> (forwards_rate*(na*cl)) : true;
[backwards] (backwards_rate*(na_plus*cl_minus)) > 0 -> (backwards_rate*(na_plus*cl_minus)) : true;
endmodule
rewards "na" true : na; endrewards
rewards "cl" true : cl; endrewards
rewards "na_plus" true : na_plus; endrewards
rewards "cl_minus" true : cl_minus; endrewards
From the latter, we can use PRISM to generate a simple plot of the expected amount of Na and Na+ over time (using both model checking and a single random trace from the simulator):
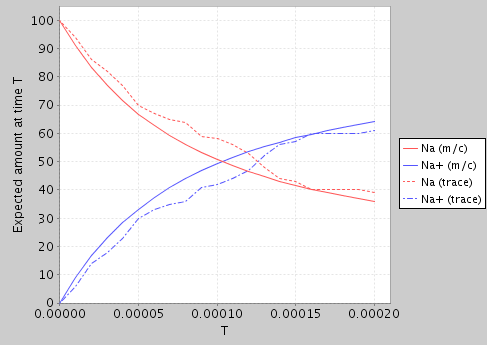 Expected amount of Na/Na+ at time T
Expected amount of Na/Na+ at time T
Using the translator
At present, the SBML-to-PRISM translator is included in the PRISM code-base, but not integrated into the application itself. To perform a conversion:
cd prism
java -cp classes prism.SBML2Prism sbml_file.xml > prism_file.sm
If you are using a binary (rather than source code) distribution of PRISM, replace classes with lib/prism.jar in the above.
Alternatively (on Linux or Mac OS X), ensure prism is in your path and then save the script below as an executable file called sbml2prism:
#!/bin/sh
# Startup script for SBML-to-PRISM translator
# Launch using main PRISM script
PRISM_MAINCLASS="prism.SBML2Prism"
export PRISM_MAINCLASS
prism "$@"
Then use:
sbml2prism sbml_file.xml > prism_file.sm
The following PRISM properties file will also be useful:
const double T;
const int c;
R{c}=? [I=T]
This contains a single property which, based on the reward structures in the PRISM model generated by the translator, means "the expected amount of species c at time T". The constant c is an integer index which can range between 1 and N, where N is the number of species in the model. To view the expected amount of each species over time, create an experiment in PRISM which varies c from 1 to N and T over the desired time range.
Details of the translation
The basic structure of the translation process is as follows:
- Each species in the SBML file is represented by a module in the resulting PRISM file. This module, which (where possible) retains the SBML species id as its name, contains a single variable whose value represents the amount of the species present. A corresponding reward structure for computing the expected amount of the species at a given time instant is also created. Species for which the
boundaryCondition flag is set to true in the SBML file do not have a corresponding module.
- Each reaction in the SBML file is associated with a unique synchronisation action label. The module for each species which takes part in the reaction will include a synchronous command to represent this. An additional PRISM module called
reaction_rates stores the expression representing the rate of each reaction (from the corresponding kineticLaw section in the SBML file). Reaction stoichiometry information is respected but must be provided in the scalar stoichiometry field of a speciesReference element, not in a separate StoichiometryMath element.
- Each parameter in the SBML file, either global to the file or specific to a reaction, becomes a constant in the PRISM file. If a value for this parameter is given, it used. If not, the constant is left as undefined.
As described above, this translation process is designed for discrete systems and so the amount of each species in the model is represented by an integer variable. It is therefore assumed that the initial amount for each species specified in the SBML file is also given as an integer. If this is not the case, then the values will need to be scaled accordingly first.
Furthermore, since PRISM is primarily a model checking (rather than simulation) tool, it is important that the amount of each species also has an upper bound (to ensure a finite state space). When model checking, the efficiency (or even feasibility) of the process is likely to be very sensitive to the upper bound(s) chosen. When using the discrete-event simulation functionality of PRISM, this is not the case and the bounds can can be set much higher. By default the translator uses an upper bound of 100 (which is increased if the initial amount exceeds this). A different value can specified through a second command-line argument as follows:
cd prism
java -cp classes prism.SBML2Prism sbml_file.xml 1000 > prism_file.sm
Alternatively, upper bounds can be modified manually after the translation process.
Finally, The following aspects of SBML files are not currently supported and are ignored during the translation process:
- compartments
- events/triggers